Calakos Lab
We all know that as part of our daily lives we are constantly interacting with our environment - learning, adapting, establishing new memories and habits, and for better or for worse, forgetting as well. At the cellular level, these processes can be encoded by changes in the strength of synaptic transmission between neurons. The process by which neuronal connections change in response to experience is known as “synaptic plasticity” and this process is a major interest of our laboratory. Our goals are to understand the molecular mechanisms for synaptic plasticity and identify when these processes have gone awry in neurological diseases. In doing so, we will establish the necessary framework to target these processes for therapeutic interventions; potentially identifying novel and improved treatment options.
We focus these interests on the striatal circuitry of the basal ganglia. The striatum is a key entry point for cortical information into the basal ganglia. The basal ganglia are involved in a wide variety of behaviors because they are critical for our movement, including the learning of motor routines and when to call them into action. Disorders in this process have wide ranging manifestations and substantially contribute to diseases like Parkinson’s disease, OCD, dystonia, Tourette’s and addictive behavior.
Our lab has pioneered a number of molecular and circuit-cracking methodologies that have provided new views into the workings of the striatal circuitry and its plasticity rules. Our lab has deep expertise in electrophysiology and optical physiology (two photon calcium imaging) and state-of-the-art molecular genetic mouse modeling techniques. Yet, our insights are further amplified by the highly collaborative approach we have with colleagues at Duke and beyond.
To get a better view of how pathway balance in basal ganglia circuitry may be affected, our lab has developed tools and approaches that make it possible to study the function of striatal medium spiny neurons in the direct and indirect pathways simultaneously in living tissue (Shuen et al., 2008; Ade et al., 2011, O’Hare and Ade et al., 2016). We use them to identify functional differences between these two types of medium spiny neurons and their role in normal adaptive plasticity and disease processes.
In habit, we identified circuit predictors of behavior. These include some classic expectations for mechanisms of plasticity such as increased firing activity, but also some surprises, like finding shifts in the timing of firing between these two cell types (O’Hare and Ade et al., 2016) and that a key coordinator is an interneuron (O’Hare et al., eLife 2017).
In disease settings, we leverage the Sapap3 KO model to understand what causes repetitive, self-injurious behavior and anxiety-like behaviors (“OCD-like”). We find a central role for striatal group 1 metabotropic glutamate receptor overactivity (Ade et al., Biol. Psych. 2016). By developing a unique high-throughput screening assay for an inherited cause of the movement disorder, dystonia, we came to recognize that multiple forms of this disease were united by a common defect in signaling by the proteostasis pathway known as the “integrated stress response” or ISR (also eIF2alpha phosphorylation) (Rittiner and Caffall et al., Neuron 2016).
Currently, ISR research in the lab has markedly expanded to address both its basic mechanisms (Helseth and Hernandez-Martinez et al., Science 2021) and its translational potential (Caffall et al., Sci. Transl. Med. 2021) for dystonia, Parkinson’s and other brain diseases.
Calakos Lab Members
Alumni
Calakos Image Gallery


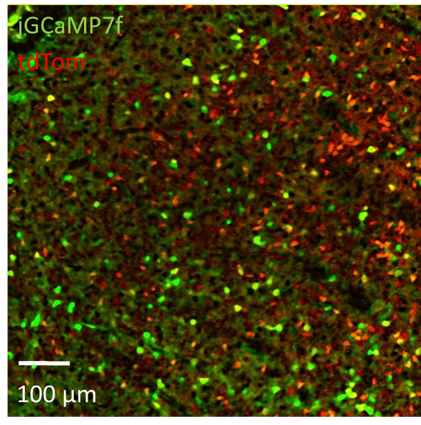
The striatum receives inputs from many areas of the brain and conveys action-related information to the basal ganglia via the direct and indirect pathways. It was long thought that an imbalance between these pathways could alter behavior, but relativistic measurements had proven elusive. By imaging activity in both pathways simultaneously, we found that relative pathway timing predicted how habitually individual mice had behaved.
In this illustration by Miranda Shipman, a bookie looks through special binoculars to pick the winner of a horse race. He sees the odds are stacked in favor of the horse, “Habit”, who has a head-start on the line, reflecting the timing bias in which firing of the direct or “Go” pathway precedes the indirect pathway, and is more muscular, reflecting the strengthening of both the direct and indirect pathways. miranda.shipman@duke.edu
Zac Caffall has designed an automated high-throughput screen to correct a cellular abnormality associated with the mutated protein that causes Dyt1 dystonia. We used it in a genome-wide silencing RNA screen. This gives us a roadmap of genes and cellular processes that modify Dyt1 cell pathology to guide us toward new treatments. Our lab is now working on a large scale drug-library screen in collaboration with the National Center for Advancing Translational Sciences (NCATS) at the NIH.

Our first 2p rig built by Kristen Ade gives us a unique view into how direct and indirect pathways of striatum are regulated in health and disease.

The 2015 lab group (and progeny and pets) chillaxing at nearby Eastwood Lake.

2015 lab outing to the U.S. National Whitewater Center.

Hipsters or posers? You be the judge – it was Halloween after all. Miranda Shipman, our “miracle maker” and resident artist and Zac Caffall, biochemist extraordinaire and all-around go-to guy.
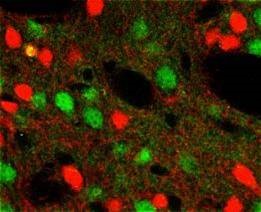
Circuit-specific physiology of striatum facilitated by transgenic reporters of cell-type. Shown here are fluorescent cells in striatum reported by our Drd1a-tdTomato line 6 (Ade et al., 2011; Shuen et al., 2008) and the Drd2-EGFP GENSAT line (Gong et al.,2003 ).

In this study, we first started thinking more about the possible role of rare variants in known dystonia for causing the more common, sporadic forms of focal dystonia (Calakos and Patel et al., 2009).
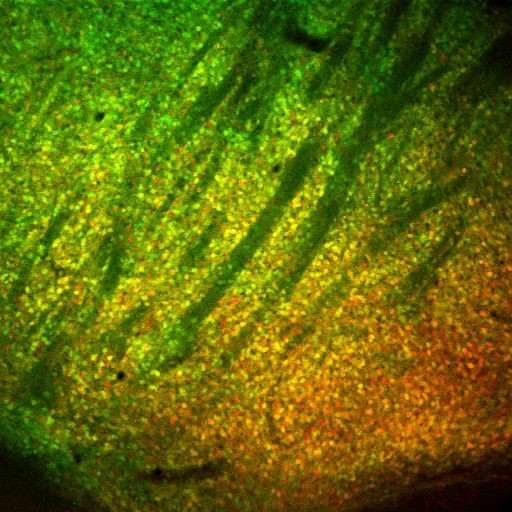
Low-mag view of striatum showing medium spiny neurons in all their glory…… (Color from Drd1a-tdTomato and Drd2-EGFP cell reporters).

Duke in Spring is a wonderful thing.

Lab alums, Yehong Wan (front) and Viren Patel (rear) floating on the Eno River just minutes from the Duke campus.
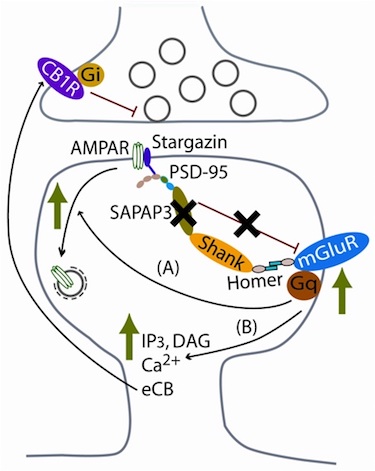
Working model for effect of SAPAP/GKAPs on regulation of mGluRs and downstream synaptic physiology.

The Calakos Lab – 2010
Calakos Open Positions
Do you have an interest in brain plasticity mechanisms? Striatal circuitry, habit, compulsion, movement disorders, dystonia, or Parkinson’s? We are always looking for a new team member to join our highly collaborative and innovative research team. If interested, please reach out to learn more about our current activities and opportunities. Send a cover letter and CV/resume to Nicole.calakos@duke.edu. We work to create and support a diverse and inclusive workspace. We welcome applicants from diverse backgrounds and starting points on their path to research and discovery excellence.
Calakos Publications
Selected Publications
Complete List of Published Work in MyBibliography
COVER: Calakos N, Zech M. Emerging Molecular-Genetic Families in Dystonia: Endosome-Autophagosome-Lysosome and Integrated Stress Response Pathways. Mov Disord. 2025
Walder-Christensen KK, Soliman HA, Calakos N, Dzirasa K. Synaptic editing of frontostriatal circuitry prevents excessive grooming in SAPAP3-deficient mice. bioRxiv [Preprint]. 2025 Mar
Bukhari-Parlakturk N, Mulcahey PJ, Lutz MW, Ghazi R, Huang Z, Dannhauer M, Termsarasab P, Scott B, Simsek ZB, Groves S, Lipp M, Fei M, Tran TK, Wood E, Beynel L, Petty C, Voyvodic JT, Appelbaum LG, Al-Khalidi HR, Davis SW, Michael AM, Peterchev AV, Calakos N. Motor network reorganization associated with rTMS-induced writing improvement in writer's cramp dystonia. Brain Stimul. 2025
Matthew L Oliver, Zachary F Caffall, Callie B Eatman, Timothy D Faw, Nicole Calakos DIO-SPOTlight Transgenic Mouse to Functionally Monitor Protein Synthesis Regulated by the Integrated Stress Response
Lockshin Elana R, Calakos N. The integrated stress response in brain diseases: A double-edged sword for proteostasis and synapses. Curr Opin Neurobiol. 2024
Calakos N, Caffall ZF. The integrated stress response pathway and neuromodulator signaling in the brain: lessons learned from dystonia. J Clin Invest. 2024
Hazlett MF, Hall VL, Patel E, Halvorsen A, Calakos N, West AE. The Perineuronal Net Protein Brevican Acts in Nucleus Accumbens Parvalbumin-Expressing Interneurons of Adult Mice to Regulate Excitatory Synaptic Inputs and Motivated Behaviors. Biol Psychiatry. 2024
Zachary F. Caffall, Bradley J. Wilkes, Ricardo Hernández-Martinez, Joseph E. Rittiner, Jennifer T. Fox, Kanny K. Wan, Miranda K. Shipman, Steven A. Titus, Ya-Qin Zhang, Samarjit Patnaik, Matthew D. Hall, Matthew B. Boxer, Min Shen, Zhuyin Li, David E. Vaillancourt, Nicole Calakos The HIV protease inhibitor, ritonavir, corrects diverse brain phenotypes across development in mouse model of DYT-TOR1A dystonia. Science Translational Medicine 18 Aug 2021:Vol. 13, Issue 607, eabd3904 DOI: 10.1126/scitranslmed.abd3904
Helseth, A.R.*, Hernandez-Martinez, R.*, Hall, V.L., Oliver, M.L., Turner, B.D., Caffall, Z.F., Rittiner, J.E., Shipman, M.K., King, C.S., Gradinaru, V., Gerfen, C., Costa-Mattioli, M., and N. Calakos. Cholinergic Neurons Constitutively Engage the ISR for Dopamine Modulation and Skill Learning. Science, 2021 Apr 23;372(6540):eabe1931.
O’Hare J, Calakos N, Yin HH. Recent Insights into Corticostriatal Circuit Mechanisms underlying Habits: Invited review for Current Opinions in Behavioral Sciences. Curr Opin Behav Sci. 2018 Apr;20:40-46.
Hernandez-Martinez, R. and N. Calakos. Seq-ing the Circuit Logic of the Basal Ganglia. Trends Neurosci, Jun, 2017.
O'Hare JK, Li H, Kim N, Gaidis E, Ade K, Beck J, Yin H, Calakos N. Striatal fast-spiking interneurons selectively modulate circuit output and are required for habitual behavior. Elife. 2017 Sep 5;6. pii: e26231. doi: 10.7554/eLife.26231.
Wang X, Gallegos DA, Pogorelov VM, O'Hare JK, Calakos N, Wetsel WC, West AE. Parvalbumin Interneurons of the Mouse Nucleus Accumbens are Required For Amphetamine-Induced Locomotor Sensitization and Conditioned Place Preference. Neuropsychopharmacology. 2017 Aug 25. doi: 10.1038/npp.2017.178. [Epub ahead of print]
Rittiner, J.E.*, Caffall, Z.F.*, Hernandez-Martinez, R., Sanderson, S.M., Pearson, J.L., Tsukayama, K.K., Liu, A.Y., Xiao, C., Tracy, S., Hickey, P., Johnson, J., Scott, B., Stacy, M., Saunders-Pullman, R., Bressman, S., Simonyan, K., Sharma, N., Ozelius, L.J., Cirulli, E.T., and N. Calakos . Functional Genomic Analyses of Mendelian and Sporadic Disease Identify Impaired eIF2α Signaling as a Generalizable Mechanism for Dystonia. Neuron, 2016 Dec 21;92(6):1238-1251.
Ade, K., Wan, Y., Hamman, H., O’Hare, J.K., Guo, W., Quian, A., Kumar, S., Bhagat, S., Rodriguiz, R.M., Wetsel, W.C., Conn, P.J., Dzirasa, K., Huber, K., and N. Calakos. 2016. Excess ongoing mGluR5 signaling drives OCD-like behavior in mice. Biological Psychiatry, May 13, PMID: 27436084.
O’Hare, J.K., Ade, K., Sukharnikova, T., Yin, H.H., and N. Calakos. Pathway-specific striatal substrates for making and breaking a habit. Neuron, 2016 (NIHMS 753257, Publ.ID: NEURON12985).
Rossi, M.A., Calakos, N., and H. H. Yin. Spotlight on Movement Disorders:What optogenetics has to offer. Movement Disorders, 2015
Deng, J., Wan, Y., Wang, X., Cohen, S., Wetsel, W.C., Greenberg, M., Kenny, P., N. Calakos, and A. West. 2014. MeCP2 phosphorylation limits psychostimulant-induced behavioral and neuronal plasticity. The Journal of Neuroscience, 34(13)4519-27.
Wan, Y., Ade, K., Caffall, Z., Ozlu, M. I., Eroglu, C., Feng, G., and N. Calakos. 2013. Circuit-selective striatal synaptic dysfunction in the Sapap3 knockout mouse model of obsessive-compulsive disorder. Biological Psychiatry.
Yang, Y. and N. Calakos. Presynaptic Long-Term Synaptic Plasticity. Front Synaptic Neurosci. 5:8. doi: 10.3389/fnsyn.2013.00008 (2013).
Farrell, M.S., Pei, Y., Wan, Y., Yadav, P., Daigle, T.L., Setola, V., Sciaky, N., Simmons, A., Nonneman, R.J., Guettier, J-M., Moy, S., Wess, J., Caron, M.G., Calakos, N., and B.L. Roth. 2012. A Gas DREADD mouse for selective modulation of cAMP production in striatopallidal neurons. Neuropsychopharmacology. 38(5):854-62.
Wan, Y., Feng, G. and N. Calakos. 2011. Sapap3 deletion causes mGluR5-dependent silencing of AMPAR synapses. The Journal of Neuroscience. 31(46):16685-91. PMC3475185.
Yang, Y. and N. Calakos. 2011. Munc13 is required for presynaptic long-term potentiation. The Journal of Neuroscience. 31(33):12053-7. PMC3201725.
Chen, M., Wan, Y., Ade, K., Ting, J., Feng, G., and N. Calakos. 2011. Sapap3 deletion causes anomalous endocannabinoid-mediated synaptic plasticity by enhancing group 1 metabotropic glutamate receptor activity. The Journal of Neuroscience, 31(26):9563-73. PMC3367431.
Ade, K., Wan, Y., Chen, M., Gloss, B. and N. Calakos. 2011. An improved BAC transgenic fluorescent reorter line for sensitive and specific identification of striatonigral medium spiny neurons. Frontiers in Neuroscience 5 (32):1-9, PMC3113108.
Yang, Y. and N. Calakos. 2010. Acute in vivo genetic rescue demonstrates that phosphorylation of RIM1 serine 413 is not required for mossy fiber LTP. The Journal of Neuroscience, 30(7):2542-6, PMC2947440.
Calakos, N., Patel, V.D., Gottron, M., Wang, G., Tran-Viet, K.-N., Brewington, D., Beyer, J.L., Steffens, D.C., Krishnan, R.R., and S. Zuchner. 2009. Functional evidence implicating a novel TOR1A mutation in idiopathic, late-onset focal dystonia. Journal of Medical Genetics, 47(9):646-50, PMC2891583.
Shuen, J.A., Chen, M., Gloss, B., and N. Calakos, 2008. Drd1a-tdTomato transgenic mice for simultaneous visualization of MSNs in the direct and indirect pathways of the basal ganglia. The Journal of Neuroscience, 28(11):2681-5.
Welch, J.M., Lu, J., Rodriguiz, R.M., Trotta, N.C., Peca, J., Ding, J-D., Feliciano, C., Adams, J.P., Dudek, S.M., Weinberg, R.J., Calakos, N., Wetsel, W.C., and Feng, G., 2007. Cortico-striatal synaptic defects and OCD-like behaviors in SAPAP3 mutant mice. Nature, 448(7156): 894-900.
Calakos, N., Schoch, S., Sudhof, T.C., and R.C. Malenka. 2004. Multiple Roles for the Active Zone Protein RIM1a in Late Stages of Neurotransmitter Release. Neuron 42(6):889-896 [Highlighted in Nat. Rev. Neurosci. 5(8):601, 2004]
Calakos, N., M.K. Bennett, K.E. Peterson and R.H. Scheller. 1994. Protein-Protein Interactions Contributing to the Specificity of Intracellular Vesicular Trafficking. Science 263:1146-1149.
Bennett, M.K., N. Calakos and R.H. Scheller. 1992. Syntaxin: A Synaptic Protein Implicated in Docking of Synaptic Vesicles at Presynaptic Active Zones. Science 257:255-259.
Bennett, M.K., N. Calakos, T. Kreiner and R.H. Scheller. 1991. Synaptic Vesicle Membrane Proteins Interact to Form a Multimeric Complex. The Journal of Cell Biology, 116:761-775.