West Lab
Epigenomic regulation of neuronal development and plasticity
Welcome to the West lab! We are a diverse and enthusiastic group of researchers investigating the molecular mechanisms of chromatin regulation that orchestrate neuronal development, synapse plasticity, and behavioral adaptations to sensory input.
The vertebrate brain is a highly adaptable organ that has a remarkable ability to convert transient environmental information into long-lasting changes in neuronal function. During development, sensory-driven neuronal activity helps to sculpt the organization of cortical circuits. In the adult brain, cell-type specific and experience-dependent plasticity of synaptic connectivity, synaptic strength, and intrinsic excitability mediate the changes in circuit function that underlie learning and memory.
Transcription factors and chromatin regulators play an essential role in these processes by coordinating the regulation of neuronal gene expression programs that are required for the formation, maturation, and plasticity of synapses. The gene expression differences that define cell type or cell state are determined by the accessibility and local conformational interactions of binding sites for these transcriptional regulators. The full set of DNA and histone marks that determine the structure of gene regulatory elements is collectively known as the chromatin landscape or the epigenome.
The dynamic nature of the epigenome is particularly intellectually compelling in the brain, because the plasticity of this biochemical process so precisely recalls the specialized functions of this organ. The persistence of epigenetic marks – which for example can commit cells to stay within a specific fate lineage for their whole lives – suggests epigenetic marks as the cellular recorder of biological experiences and a mechanism of neuronal memory. Yet another fundamental physiological property of the epigenome is stimulus-inducible reversibility, thus providing cells with plasticity to adapt in response to changes in their environment, which echoes the principle requirements for a mechanism of learning.
Brain plasticity confers an evolutionary advantage by allowing organisms to adapt their behavior to the environment. However aberrant plasticity can also predispose the vertebrate brain to neurological and neuropsychiatric illnesses. Genetic mutations in transcription factors and chromatin regulators are associated with neurodevelopmental disorders and cognitive impairments, demonstrating the biological importance of gene regulation in the human brain. Dysregulation of transcriptional plasticity is associated with neuropsychiatric disorders including addiction and depression.
We seek to discover the chromatin mechanisms that mediate experience-dependent brain development and plasticity both to understand at a fundamental level how the brain adapts to its environment and to contribute to strategies that can correct maladaptive changes in brain development and function.
West Lab Projects
We use molecular genetics, experimental model systems, sophisticated behavioral analyses, biochemical and cell biological methods, and high-throughput sequencing techniques to elucidate transcriptional mechanisms that underlie neuronal adaptations to changes in the environment. In addition to enhancing understanding of normal brain development and plasticity, our studies are revealing new insights into how dysregulation of gene transcription and chromatin regulation contributes to neurodevelopmental disorders and mental illness.
Click on the + signs below to read more about current projects in the lab.
Cellular differentiation requires the precise spatial and temporal orchestration of gene expression programs. Progressive changes in gene transcription during development are driven by epigenetic modifications of genomic DNA and its associated histone proteins, collectively called chromatin, that differentially alter the access of DNA regulatory sequences to the transcriptional machinery. A growing body of evidence shows that chromatin-regulatory proteins can be modulated by environmental stimuli, raising the intriguing possibility that early life experience may impact development by inducing plasticity of the epigenome. Neurons are extremely long-lived cells that, as part of their fundamental function in information processing, undergo robust and dynamic changes in their gene expression repertoires long after they have left the cell cycle and committed to a postmitotic identity. Thus neurons are ideal for studying functions of the dynamic epigenome beyond its role in establishing cell identity.
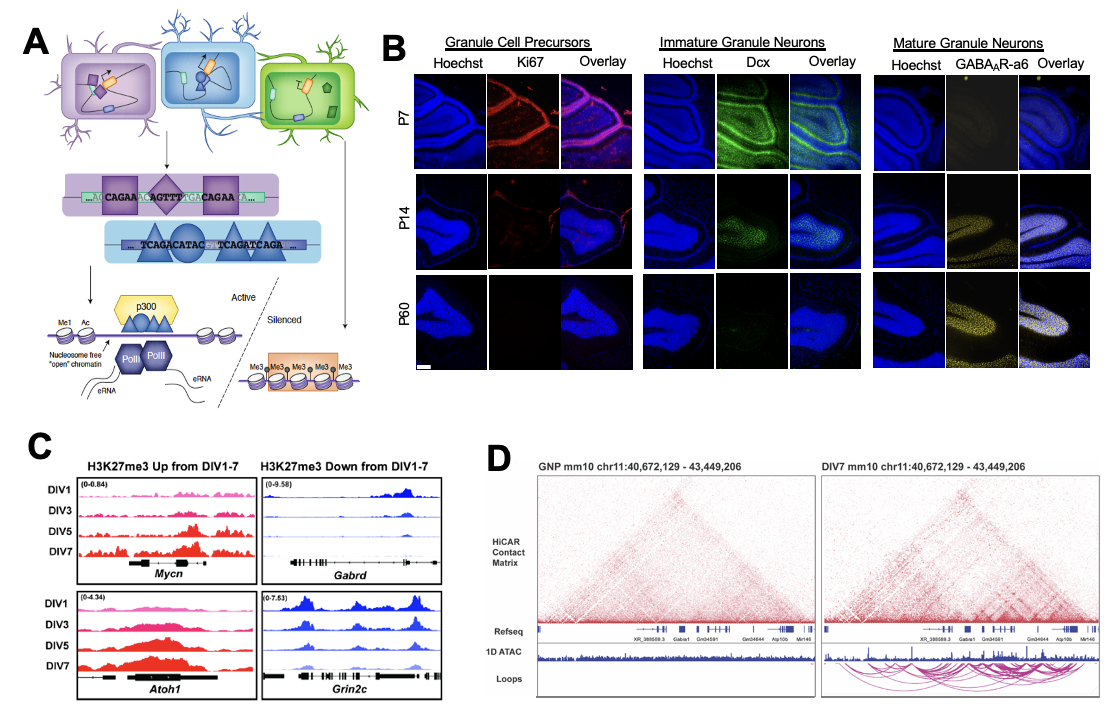
We have used the differentiation of cerebellar granule neurons (CGNs) in the postnatal brains of male and female mice as a model system for identifying molecular mechanisms of neuronal maturation. The huge abundance and postnatal development of CGNs makes them an ideal cell type for biochemical studies of the molecular mechanisms that guide neuronal differentiation. CGNs account for >99% of all neurons and >85% of all cells in the cerebellar cortex, lowering the barriers to studying a specific cell type developing in vivo. CGNs derive from committed granule neuron precursors (GNPs). Upon leaving the cell cycle these progenitors develop first into immature neurons and then into mature neurons, which can be distinguished by their unique gene expression programs. By comparing multiple stages of CGN differentiation we can identify chromatin mechanisms of gene transcription that operate during differentiation and maturation both in proliferating and in postmitotic cells of the neuronal lineage.
We are using state of the art approaches in chromatin profiling including ATAC-seq, CUT&RUN, CUT&Tag, and the low-input chromatin conformation protocol HiCAR to build a foundational understanding of the regulatory logic of dynamic chromatin states that underlie stages of neuronal maturation. We can also track the morphological development and circuit integration of newly born CGNs in vivo using in vivo electroporation of granule neuron progenitors to obtain sparse labeling and cell-autonomous genetic manipulation of these neurons.
Chromatin-regulatory proteins are among the most common classes of genes mutated in autism spectrum disorder (ASD) and intellectual disability (ID). It is well established that cellular differentiation requires the precise orchestration of gene expression programs, which is mediated by dynamic changes to chromatin state and structure. However, it remains unclear why de novo mutations in a subset of chromatin regulators give rise to the specific neurological disruptions that underlie ASD/ID. Synaptic proteins are the other major class of gene products that show de novo mutations in ASD/ID, and disruption of neuronal connectivity is thought to be a major underlying disease pathology in ASD. Our overarching hypothesis is that there are convergent roles played by multiple ASD/ID-associated chromatin regulators that mediate the orchestrated activation of genes underlying synapse development in postmitotic neurons. Our focus on neurons in the postnatal brain is relevant to therapeutics because by the time ASD/ID is diagnosed in early childhood, there is little that can be done to change fetal development, whereas gene therapy has the potential to correct ongoing deficiencies in neuronal function.
We focus in detail on the biochemical and cellular mechanisms by which ASD/ID-associated mutations in specific chromatin regulators result in impaired neuronal development.
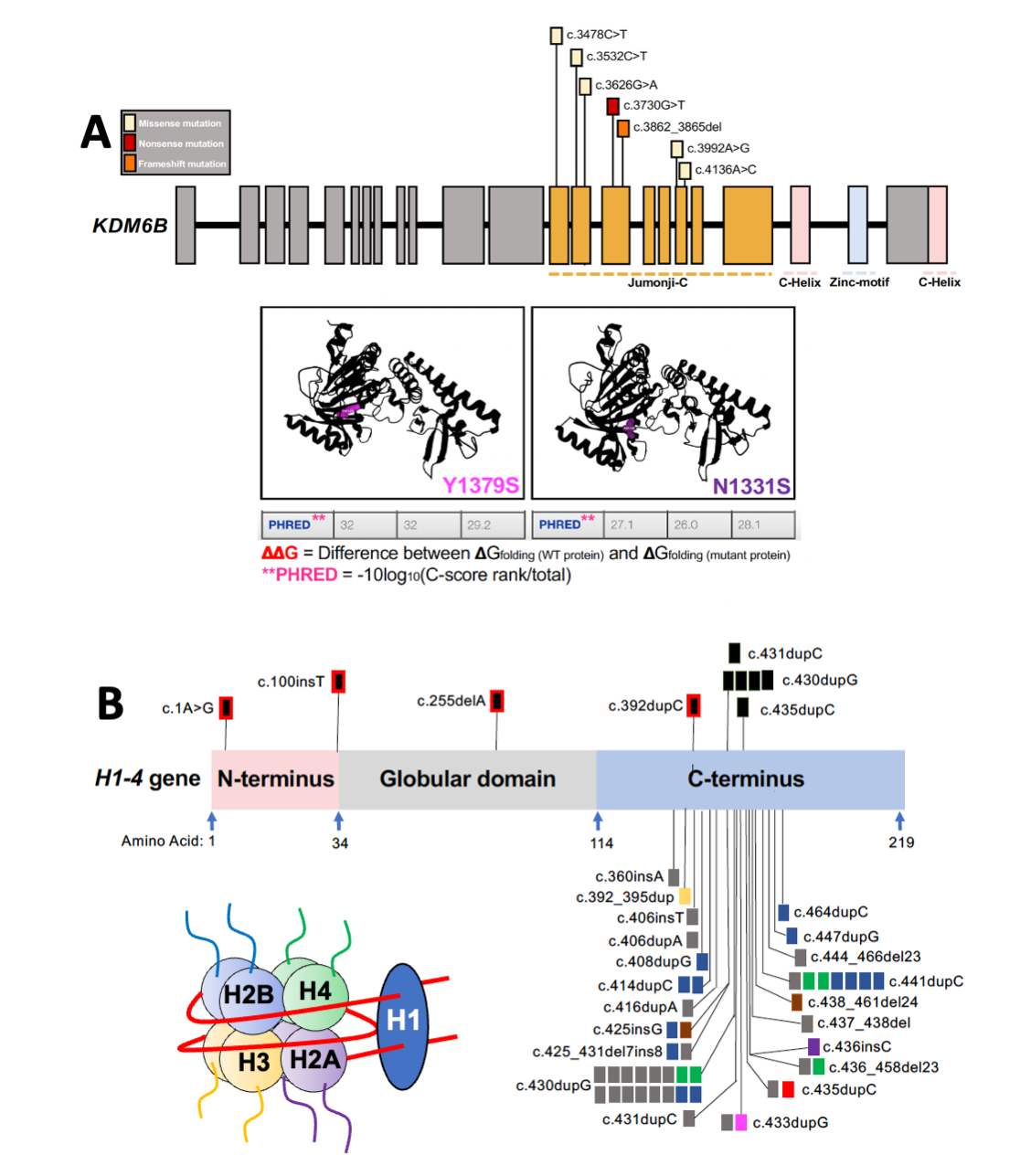
One project examines functions of the lysine demethylase Kdm6b, which remodels the distribution of the repressive histone modification H3K27me3 in developing neurons. We have shown that disruption of the lysine demethylase function of KDM6B impairs synaptic gene expression and synapse development. This is notable given evidence of ASD/ID-associated mutations in KDM6B, including point mutations in the lysine demethylase region.
A second project investigates the biochemical and cellular mechanisms by which ID-associated mutations in the variant linker histone H1.4 impair neuronal development. Histone H1 proteins bind at the entry and exit sites of nucleosomes and contribute to chromatin compaction and 3D architecture. Frame-shift mutations that disrupt the C-terminal of H1.4 cause Rahman Syndrome, a genetic form of ID. We have shown that expression of mutant H1.4 expression in neurons disrupts synaptic gene expression and are working to establish the mechanism.
In the brain, synaptic activity provides a salient stimulus for the adaptation of neuronal function through the induction of stimulus-dependent gene transcription. Distal transcriptional enhancers play both permissive and instructive roles in this process. In their permissive role, the pattern of active enhancers predetermines which genes can be induced in any given kind of neuron, such that distinct programs of activity-regulated gene expression are induced in different types of neurons, even in response to a common stimulus. A functionally important corollary is that different types of neurons adapt in distinct ways to changes in neuronal firing. At the instructive level, enhancers serve as docking sites for activity-inducible transcription factors, thus conferring temporal regulatory activity upon a given gene.
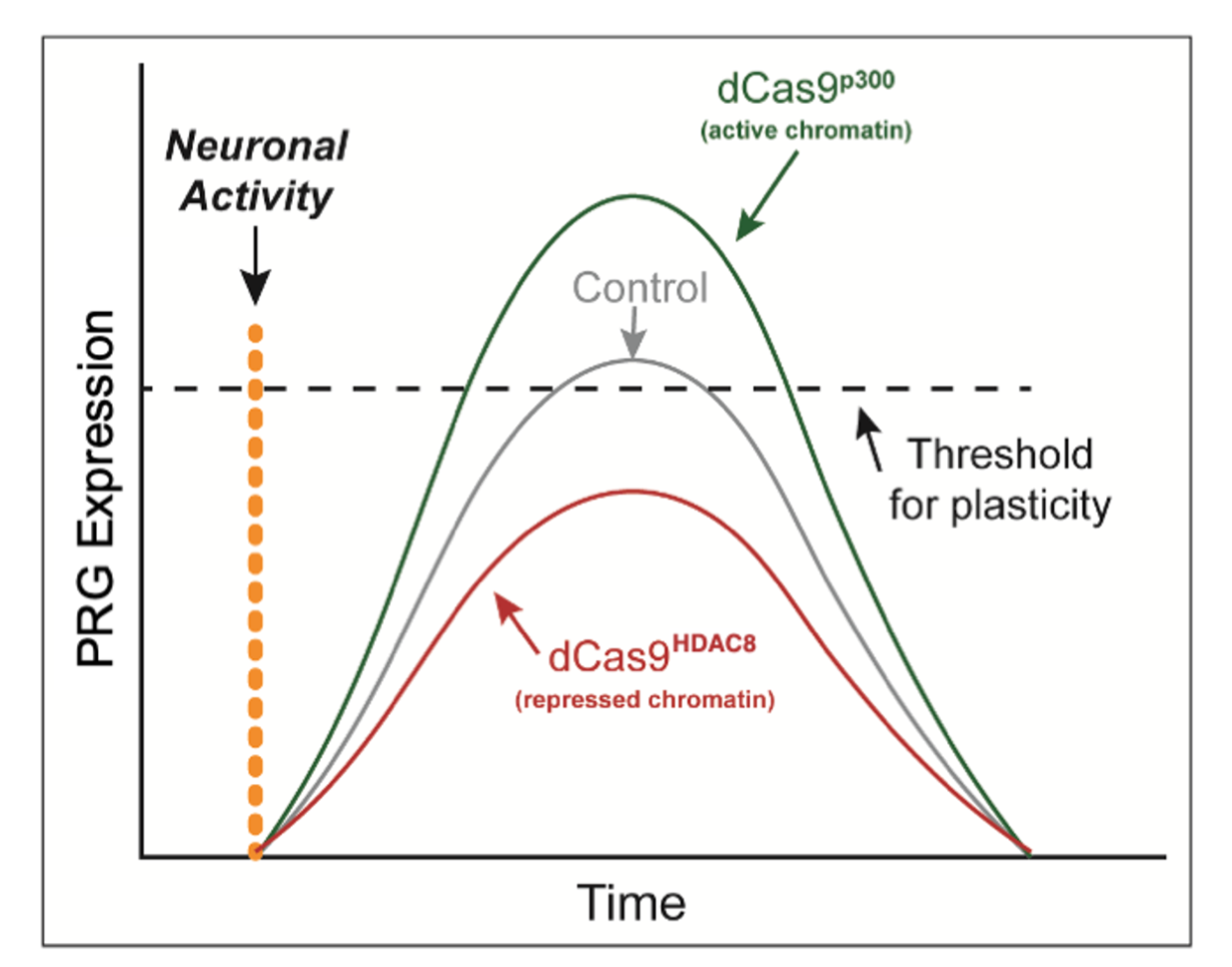
We have been studying the importance of enhancers in the context of transcriptional responses by drugs of abuse that contribute to the development of addiction. Chronic cocaine abuse arises as a result of persistent cocaine-induced adaptations in the function of the neurons that comprise mesolimbocortical brain reward circuits. Cocaine-induced changes in gene transcription contribute to many of these alterations in neuronal function. Furthermore cocaine exposure has been shown to dynamically alter the epigenome by regulating the expression and/or function of histone and DNA modifying enzymes. Taken together, these data have led to the hypothesis that long-lasting changes in the epigenome, especially at enhancers, may underlie the persistence of cocaine-induced addictive-like behaviors.
We are testing this hypothesis by using a dCas9/CRISPR based, functional genomics strategy to modify the regulatory state of the enhancer elements of two foundational activity-regulated genes: the neurotrophin Bdnf and the synaptic plasticity gene Arc. We hypothesize that regulating the epigenome at these enhancers will titrate the quantity of activity-induced gene expression to set thresholds for the induction of reward-driven motivated behaviors.
We seek to discover mechanisms that link genome regulation with neuronal and behavioral plasticity. To achieve this goal, we deploy a broad range of experimental approaches including, but not limited to, chromatin sequencing, single molecule fluorescence in situ hybridization, bioinformatic modeling, mouse genetics, and behavior. We also take advantage of novel leading-edge methods that can be applied in new ways to advance understanding of our system. For example we have combined mice carrying cell-type specific transgenes with single nuclear RNA sequencing to identify gene regulatory programs within rare interneuron populations that reside in physiologically relevant neural circuits of behaving mice.
Ongoing projects in the lab are pioneering the use of multiple new technologies in neurons including the following:
1) To understand the three-dimensional organization of genomic DNA in neurons, we are using a novel low-input chromatin sequencing method called HiCAR that was developed by our colleague Yarui Diao in the department of Cell Biology. Because HiCAR is more efficient than standard HiC, we have been able to assess the dynamics of genome architecture changes in neurons across developmental stages, in response to drug treatments that disrupt epigenome features, and in genetic models of neurodevelopmental disorders. See data in project on chromatin regulation of postmitotic neuronal maturation.
2) To experimentally edit the epigenome and then determine the consequences for neuronal and network function in vivo, we are using dCas9-CRISPR based mouse strains engineered by our colleague Charlie Gersbach in the department of Biomedical Engineering. We are also developing new dCas9 fusions to expand the ways that we can locally modify transcription along the genome. See Gemberling et al. (2021) Nat Methods 18: 965-74.
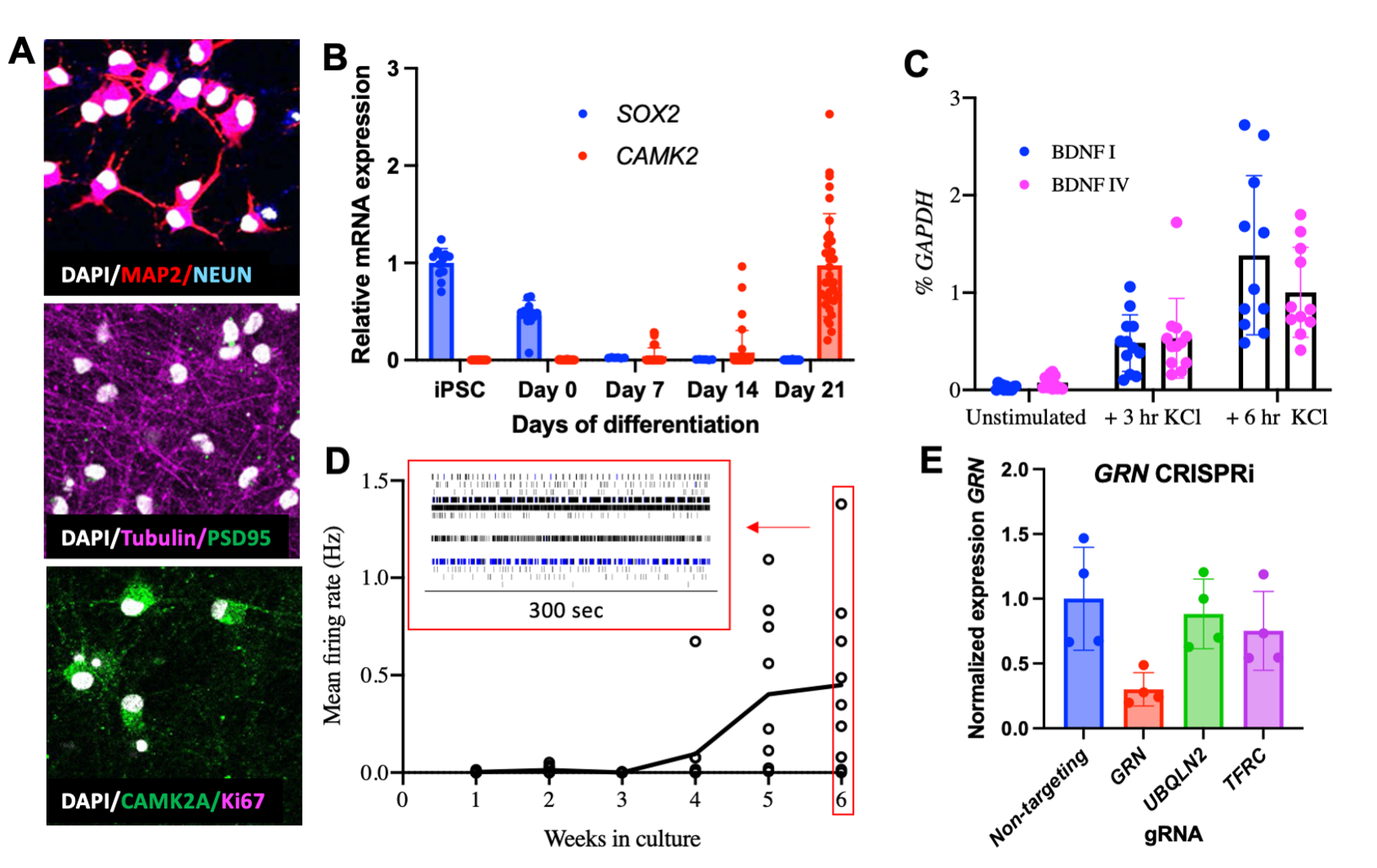
3) To study the function of chromatin regulators in the context of the human genome, we are expanding our experimental systems to include NGN2-induced human iPSC-derived neurons. The ease of generating large numbers of relatively homogenous neurons from genetically manipulatable iPSCs makes these cells ideal for studying the functional consequences of ASD/ID-associated gene mutations using sequencing assays and molecular genetic screens. We are also using these cells to study human-specific gene regulatory elements.
West Lab Members
West Lab Photos
















West Publications (Selected)
Ramesh, Vijyendra, Fang Liu, Melyssa S. Minto, Urann Chan, and Anne E. West. “Bidirectional regulation of postmitotic H3K27me3 distributions underlie cerebellar granule neuron maturation dynamics.” Elife 12 (April 24, 2023). https://doi.org/10.7554/eLife.86273.
Green, M. V., D. A. Gallegos, J. V. Boua, L. C. Bartelt, A. Narayanan, and A. E. West. “Single-Nucleus Transcriptional Profiling of GAD2-Positive Neurons From Mouse Lateral Habenula Reveals Distinct Expression of Neurotransmission- and Depression-Related Genes.” Biological Psychiatry Global Open Science, January 1, 2023. https://doi.org/10.1016/j.bpsgos.2023.04.004.
Tremblay, Martine W., Matthew V. Green, Benjamin M. Goldstein, Andrew I. Aldridge, Jill A. Rosenfeld, Haley Streff, Wendy D. Tan, et al. “Mutations of the histone linker H1-4 in neurodevelopmental disorders and functional characterization of neurons expressing C-terminus frameshift mutant H1.4.” Hum Mol Genet 31, no. 9 (May 4, 2022): 1430–42. https://doi.org/10.1093/hmg/ddab321.
Gallegos, David A., Melyssa Minto, Fang Liu, Mariah F. Hazlett, S. Aryana Yousefzadeh, Luke C. Bartelt, and Anne E. West. “Cell-type specific transcriptional adaptations of nucleus accumbens interneurons to amphetamine.” Mol Psychiatry, February 16, 2022. https://doi.org/10.1038/s41380-022-01466-1.
Gemberling, Matthew P., Keith Siklenka, Erica Rodriguez, Katherine R. Tonn-Eisinger, Alejandro Barrera, Fang Liu, Ariel Kantor, et al. “Transgenic mice for in vivo epigenome editing with CRISPR-based systems.” Nat Methods 18, no. 8 (August 2021): 965–74. https://doi.org/10.1038/s41592-021-01207-2.
Chan, Urann, Diwas Gautam, and Anne E. West. “Utilizing In Vivo Postnatal Electroporation to Study Cerebellar Granule Neuron Morphology and Synapse Development.” J Vis Exp, no. 172 (June 9, 2021). https://doi.org/10.3791/62568.
Green, Matthew V., and Anne E. West. “TRPing into excitotoxic neuronal death.” Cell Calcium 93 (January 2021): 102331. https://doi.org/10.1016/j.ceca.2020.102331.
Chen, Liang-Fu, Michelle R. Lyons, Fang Liu, Matthew V. Green, Nathan G. Hedrick, Ashley B. Williams, Arthy Narayanan, Ryohei Yasuda, and Anne E. West. “The NMDA receptor subunit GluN3A regulates synaptic activity-induced and myocyte enhancer factor 2C (MEF2C)-dependent transcription.” J Biol Chem 295, no. 25 (June 19, 2020): 8613–27. https://doi.org/10.1074/jbc.RA119.010266.
Nord, Alex S., and Anne E. West. “Neurobiological functions of transcriptional enhancers.” Nat Neurosci 23, no. 1 (January 2020): 5–14. https://doi.org/10.1038/s41593-019-0538-5.
Swahari, Vijay, and Anne E. West. “Histone demethylases in neuronal differentiation, plasticity, and disease.” Curr Opin Neurobiol 59 (December 2019): 9–15. https://doi.org/10.1016/j.conb.2019.02.009.
West, Anne E. “Activity-Dependent Transcription Collaborates with Local Dendritic Translation to Encode Stimulus-Specificity in the Genome Binding of NPAS4.” Neuron 104, no. 4 (November 20, 2019): 634–36. https://doi.org/10.1016/j.neuron.2019.10.022.
Tonn Eisinger, Katherine R., and Anne E. West. “Transcribing Memories in Genome Architecture.” Trends Neurosci 42, no. 9 (September 2019): 565–66. https://doi.org/10.1016/j.tins.2019.06.002.
Contact/Join West Lab
West Lab Mission Statement
To reveal something new about the brain,
By answering rigorously defined questions,
And to enjoy the process of discovery.
We are always looking for talented and motivated graduate students and postdocs to join our team. Undergraduate researcher and technician positions are often available. Send your CV and a brief description of your research interests to the email address below.
Anne E. West, M.D., Ph.D.
Professor
Department of Neurobiology
Email: west@neuro.duke.edu
Phone: 919-681-1909
Location: 301D Bryan Research




