Naumann Lab
The Naumann Lab is interested in how neural circuits across the entire brain guide behavior and how individuality manifests within these circuits. Our goal is to understand how precisely the neural circuits underlying sensory systems of individuals form, develop, and function. To dissect such circuits, we use the genetically accessible and translucent zebrafish to map, monitor, and manipulate neuronal activity. By combining whole-brain imaging, behavioral monitoring, functional perturbations, neuroanatomy, computational modeling, and machine learning, we aim to generate brain-scale circuit models of simple behaviors in individual brains.
Naumann Research
Optomotor response idiosyncrasy
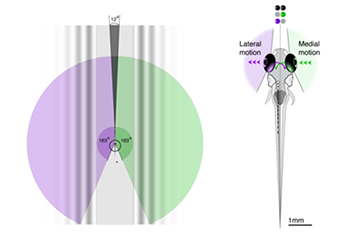
Neural circuit models that motivate specific connectivity based on average data may lead to over-interpretations in which best-fit models do not represent real neural implementations in individual animals. We are interested in how visually evoked behavioral idiosyncrasies manifest in underlying neural circuit architectures.
Brain-body interactions
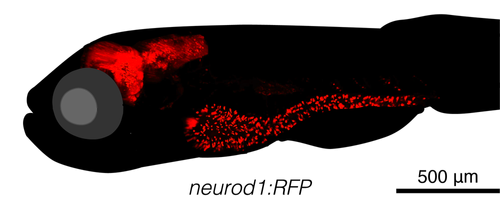
Interoceptive signals are increasingly recognized to impact behavior. We aim to dissect the molecular and functional mechanisms of brain-body communication, specifically the gut-brain axis.
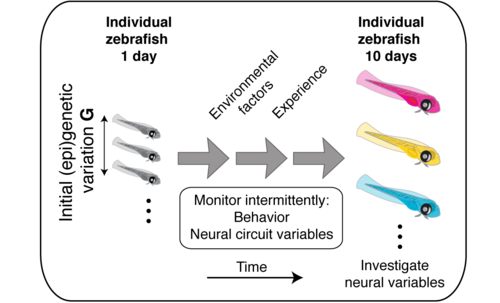
Lifetime
With longitudinal assays of behavior and neural activity, we plan to build a better understanding of how genotype and environment interact to shape neural circuits and behavior.
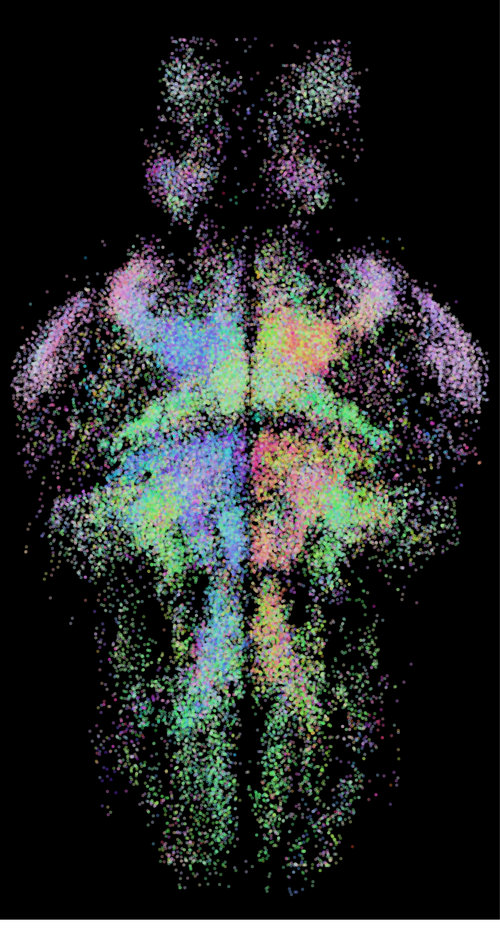
Motivational states
Motivational states, such as arousal, sleep, fear, and hunger, affect stimulus sensitivity and influence behavior. We aim to reveal how neuromodulators can flexibly alter activity in well-defined neural circuits to affect behavior and behavioral idiosyncrasy.
Lab Members
Naumann Publications
Loring, Matthew D., Eric E. Thomson, and Eva A. Naumann. “Whole-brain interactions underlying zebrafish behavior.” Curr Opin Neurobiol 65 (December 2020): 88–99. https://doi.org/10.1016/j.conb.2020.09.011.
Harfouche, M., T. W. Dunn, E. A. Naumann, and R. Horstmeyer. “Imaging the behavior and neural activity of freely moving organisms with a gigapixel microscope.” In Optics Infobase Conference Papers, Vol. Part F169-BRAIN 2019, 2019. https://doi.org/10.1364/BRAIN.2019.BT3A.3.
Naumann, Eva A., James E. Fitzgerald, Timothy W. Dunn, Jason Rihel, Haim Sompolinsky, and Florian Engert. “From Whole-Brain Data to Functional Circuit Models: The Zebrafish Optomotor Response.” Cell 167, no. 4 (November 3, 2016): 947-960.e20. https://doi.org/10.1016/j.cell.2016.10.019.
Dunn, Timothy W., Yu Mu, Sujatha Narayan, Owen Randlett, Eva A. Naumann, Chao-Tsung Yang, Alexander F. Schier, Jeremy Freeman, Florian Engert, and Misha B. Ahrens. “Brain-wide mapping of neural activity controlling zebrafish exploratory locomotion.” Elife 5 (March 22, 2016): e12741. https://doi.org/10.7554/eLife.12741.
Dunn, Timothy W., Christoph Gebhardt, Eva A. Naumann, Clemens Riegler, Misha B. Ahrens, Florian Engert, and Filippo Del Bene. “Neural Circuits Underlying Visually Evoked Escapes in Larval Zebrafish.” Neuron 89, no. 3 (February 3, 2016): 613–28. https://doi.org/10.1016/j.neuron.2015.12.021.
Randlett, Owen, Caroline L. Wee, Eva A. Naumann, Onyeka Nnaemeka, David Schoppik, James E. Fitzgerald, Ruben Portugues, et al. “Whole-brain activity mapping onto a zebrafish brain atlas.” Nat Methods 12, no. 11 (November 2015): 1039–46. https://doi.org/10.1038/nmeth.3581.
Naumann, Eva A., Adam R. Kampff, David A. Prober, Alexander F. Schier, and Florian Engert. “Monitoring neural activity with bioluminescence during natural behavior.” Nat Neurosci 13, no. 4 (April 2010): 513–20. https://doi.org/10.1038/nn.2518.
Naumann, E. A. “Illuminating the neural circuitry underlying larval zebrafish behavior,” n.d.








